Introduction:
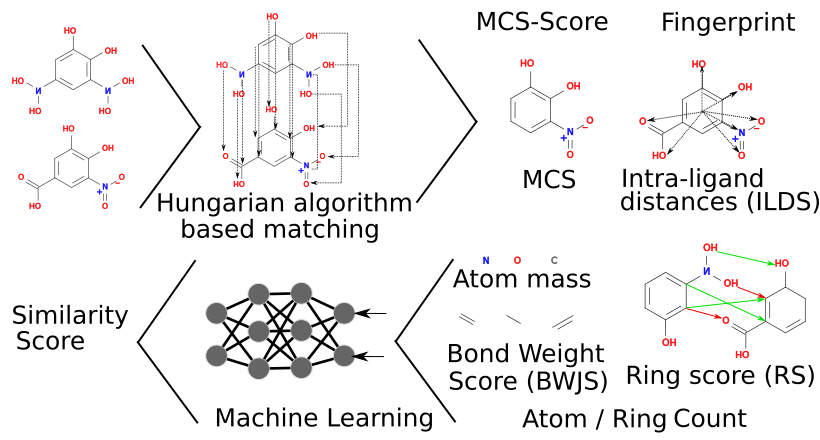
Ligand-similarity-based virtual screening is one of the most applicable computer-aided drug design techniques. Current methodology relies heavily on several descriptors of molecular features including atoms (0D), the presence or absence of structural features (1D), topological descriptors (2D), geometry and volume (3D), or stereoelectronic and stereodynamic properties (4D). These descriptors have been frequently used in virtual screening; however, they are usually used independently without integration, which may hinder effective and precise virtual screening. In this study, we developed a multi-feature integration algorithm named LigMate, which employs a Hungarian algorithm-based matching and a machine learning-based non-linear combination of various descriptors, including the new relevant descriptors focusing on the maximum common substructures (MCSS), the relative distance of atoms from the ligand mass center (ILDS), as well as the ring differences (RS). In benchmark tests, LigMate has an overall enrichment factor of the first percent (EF1) of 36.14 and an area under the curve value (AUC) of 0.81 on the DUD-E data set as well as an EF1 of 15.44 and an AUC of 0.69 on the MUV data set, which outperforms other well-established single-descriptor-based methods. Thus, our study suggests a new framework of multiple feature integration, which can be beneficial for ligand-similarity-based virtual screening
Please cite "LigMate: a Multi-Feature Integration Algorithm for Ligand-Similarity-Based Virtual Screening. Maximilian Grimm, Yang Liu, Xiaocong Yang, Bing Li, Chunya Bu, Zhixiong Xiao, Yang Cao. , 2020"
LigMate is free for non-commercial users. Other users please contact cy_scu@yeah.net .